Latest News

Uncovering Hidden Authors with AI
USC ISI computer scientists are developing language technologies that could potentially identify who wrote anonymous text.
ISI Researchers Pioneer Developments In Protein Data Bank
ISI researchers are part of a core team developing the PDB-Dev repository, initiated by the Protein Data Bank, a foundational repository for macromolecular structures
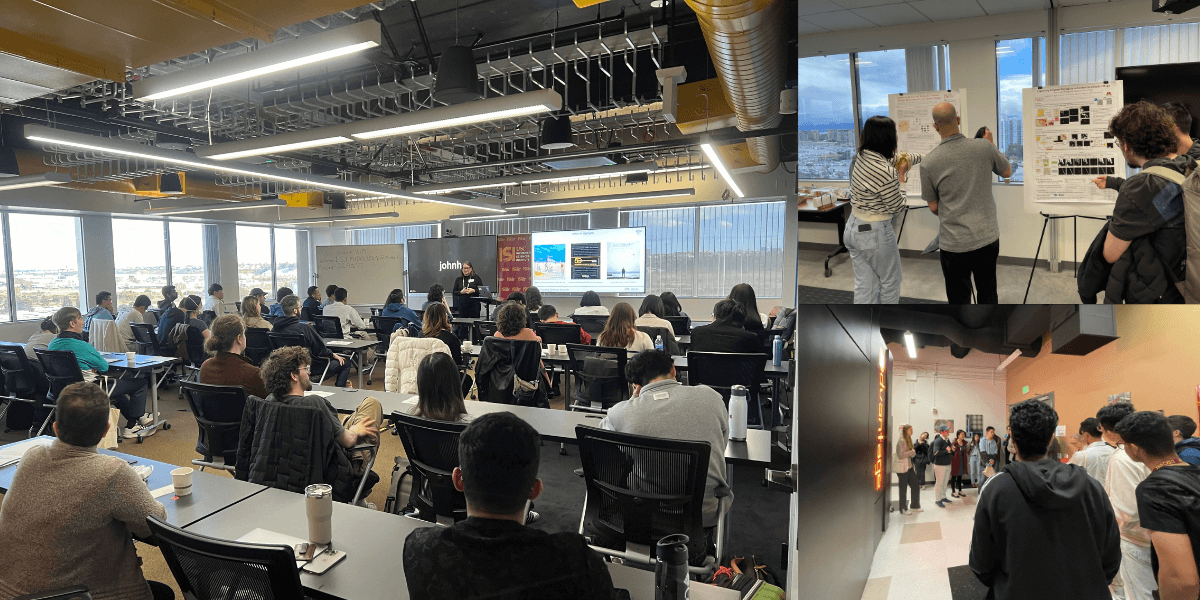

Inside ISI’s Ph.D. Visit Day 2024
Prospective graduate students traveled to ISI’s Marina Del Rey headquarters to experience the institute’s academic environment.
UPCOMING EVENTS
Apr
25
NL Seminar- How to Steal ChatGPT’s Embedding Size, and Other Low-rank Logit Tricks
Matthew Finlayson, USC
-
Apr
26
AI Seminar: Staring into the Abyss and Eating Glass
Rajiv Maheswaran, Second Spectrum
-
May
3
AI Seminar: Understanding LLMs through their Generative Behavior, Successes and Shortcomings
Swabha Swayamdipta, USC
-
