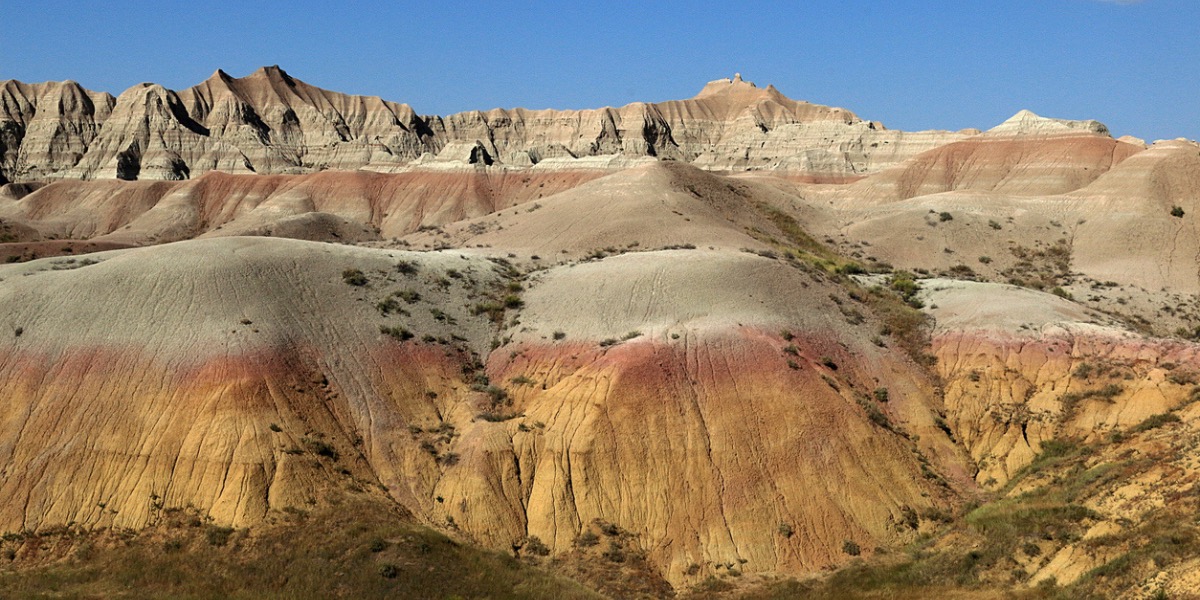
Bridging Disciplines: The Power of Collaboration at ISI
ISI connects researchers across USC to ask better questions and build smarter solutions.

Carl Kesselman Honored with 2023 IEEE Internet Award
The Information Sciences Institute Fellow will receive the prestigious award.
USC Computer Scientists Are Tackling Dental Health and Birth Defects
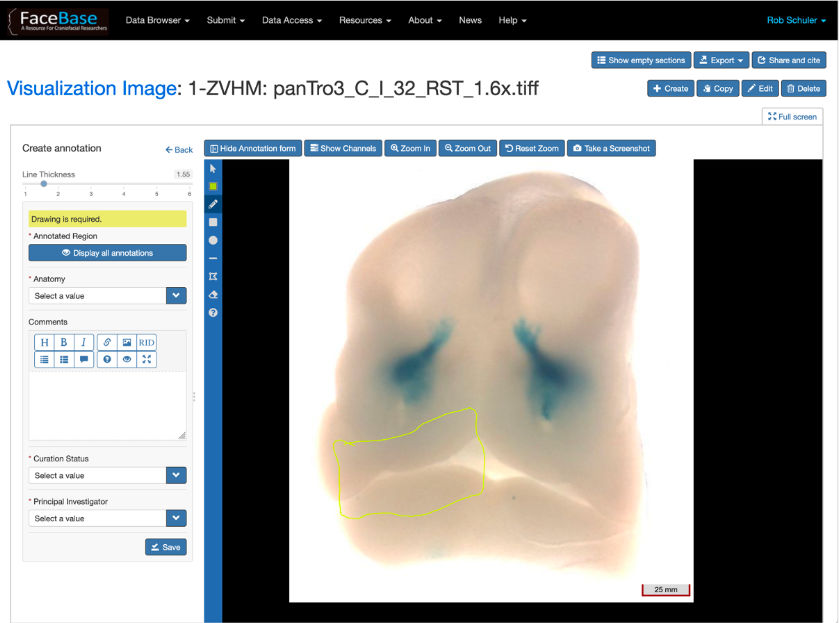
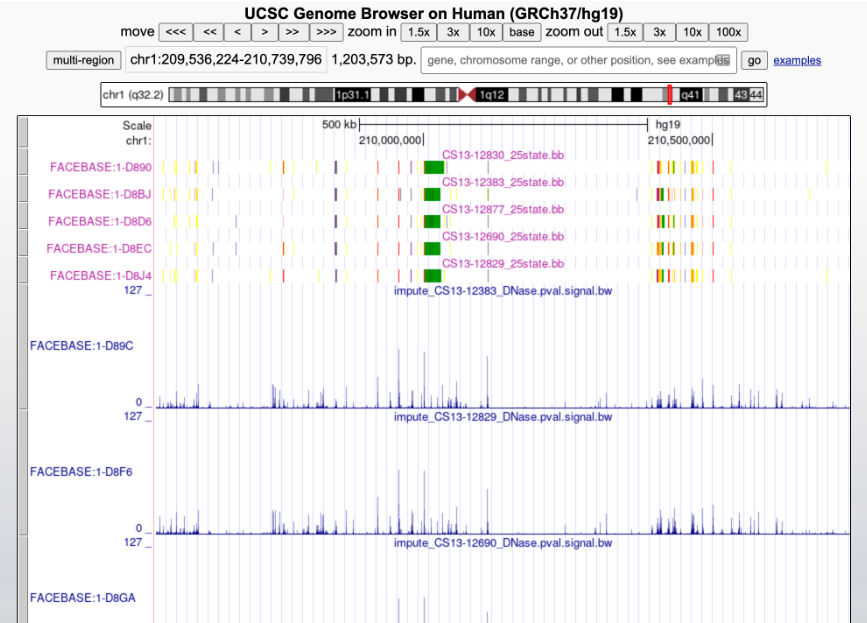
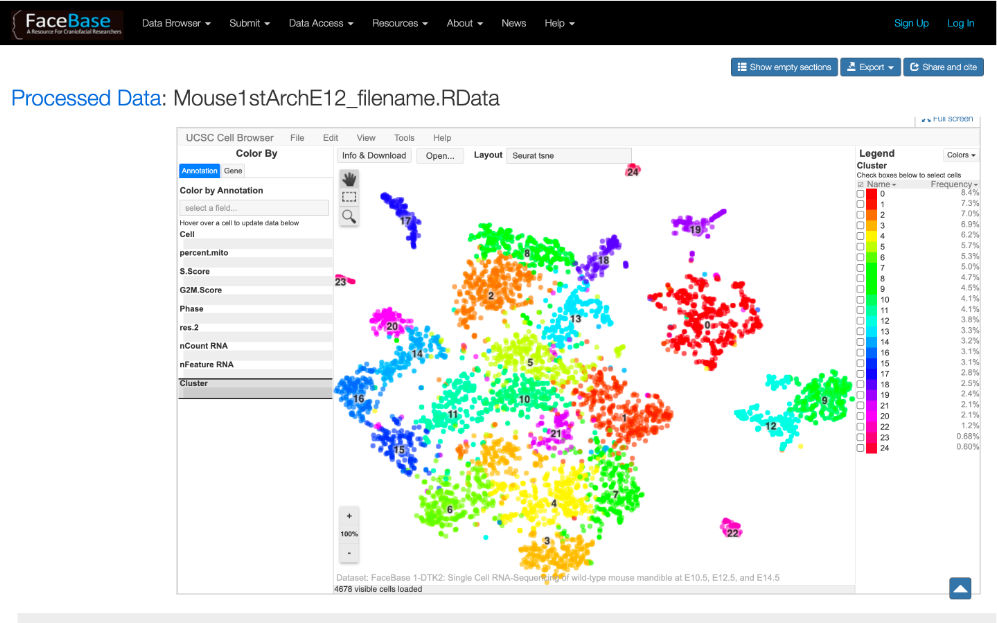
FaceBase researchers are advancing dental, oral and craniofacial research with data.
FaceBase and fair: a review
The FaceBase Consortium, funded by the National Institute of Dental and Craniofacial Research of the National Institutes of Health, was established in 2009 with the recognition that dental and craniofacial research are increasingly data-intensive disciplines. Data sharing is critical for the validation and reproducibility of results as well as to enable reuse of data. In service of these goals, data ought to be FAIR: Findable, Accessible, Interoperable, and Reusable.
The FaceBase data repository and educational resources exemplify the FAIR principles and support a broad user community including researchers in craniofacial development, molecular genetics, and genomics. FaceBase demonstrates that a model in which researchers “self-curate” their data can be successful and scalable. We present the results of the first 2.5 years of FaceBase’s operations as an open community and summarize the data sets published during this period. We then describe a research highlight from work on the identification of regulatory networks and noncoding RNAs involved in cleft lip with/without cleft palate that both used and in turn contributed new findings to publicly available FaceBase resources.
Collectively, FaceBase serves as a dynamic and continuously evolving resource to facilitate data-intensive research, enhance data reproducibility, and perform deep phenotyping across multiple species in dental and craniofacial research.
ISR members who helm the FaceBase Hub just published a review article in a special issue of the Journal of Dental Research (Data-Driven Analytics for Dental, Oral, and Craniofacial Health Care) titled “FaceBase: A Community-Driven Hub for Data-Intensive Research”. This review article summarizes the last two and a half years of FaceBase’s community-driven, self-curation model and describes the resulting varied datasets published as a result. It also describes the importance of FAIR principles for data sharing.
FaceBase: A Community-Driven Hub for Data-Intensive Research
Scientific Projects
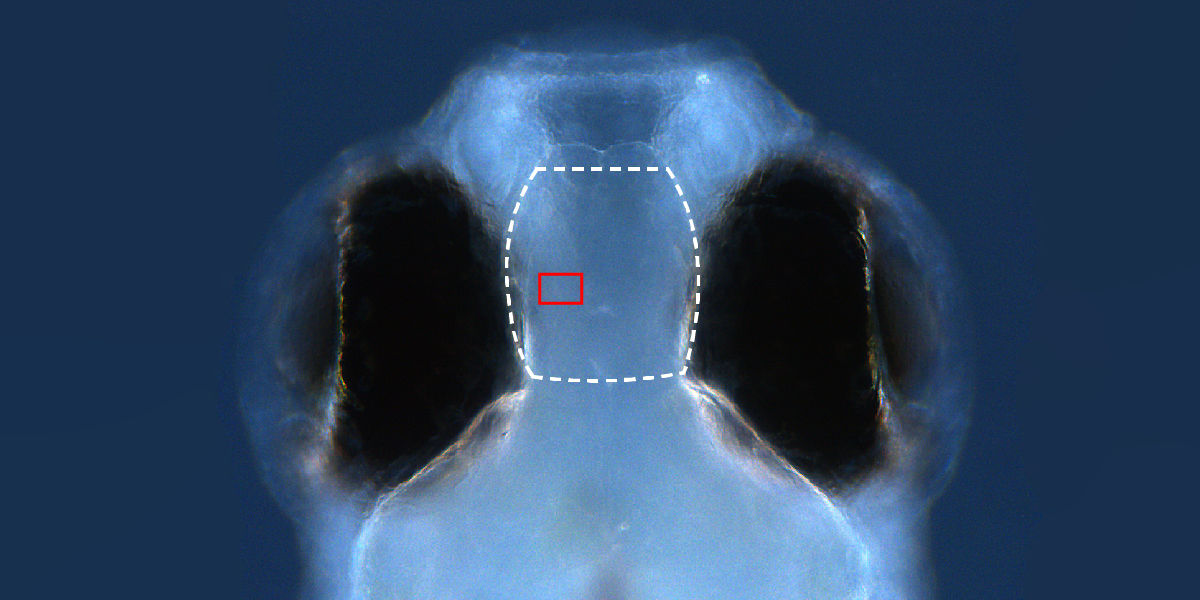
Synapse
We have developed and applied a set of tools that enables direct observation and spatial mapping of synapses using high-contrast selective plane illumination imaging of an endogenous postsynaptic density protein labeled with a recombinant GFP-coupled probe. Using these tools, we have observed changes in the microstructure of the pallium of larval zebrafish that occur during memory formation as a result of exposure to a novel classical conditioning paradigm that we developed. Surprisingly, we found that memory formation due to classical conditioning is associated with generation of new synapses, but not with systematic changes in the strengths of existing synapses.
Protein Data Bank
We are developing the necessary infrastructure to make the integrative structures of biomolecular systems available from the Protein Data Bank (PDB). PDB is the single global repository for three-dimensional structures of biological macromolecules and their complexes. It is used by millions of scientists and educators worldwide. This project will provide open access to integrative structures through the PDB, which will greatly enhance the understanding of cellular and molecular biology and speed the progress of biomedical and biotechnology research.
Learn More

Common Fund Data Ecosystem (CFDE)
NIH Data Commons, a shared virtual space where scientists can work with the digital objects of biomedical research making data-gathering more effective, reliable and accessible The CFDE is an online portal being developed to allow researchers access data from multiple programs in the cloud. ISR is working with CFDE to leverage DERIVA, a cloud-based system they have developed, to help scientists manage big scientific data as easily as pictures on a cell phone.
Learn More
Coordinating Centers
FaceBase
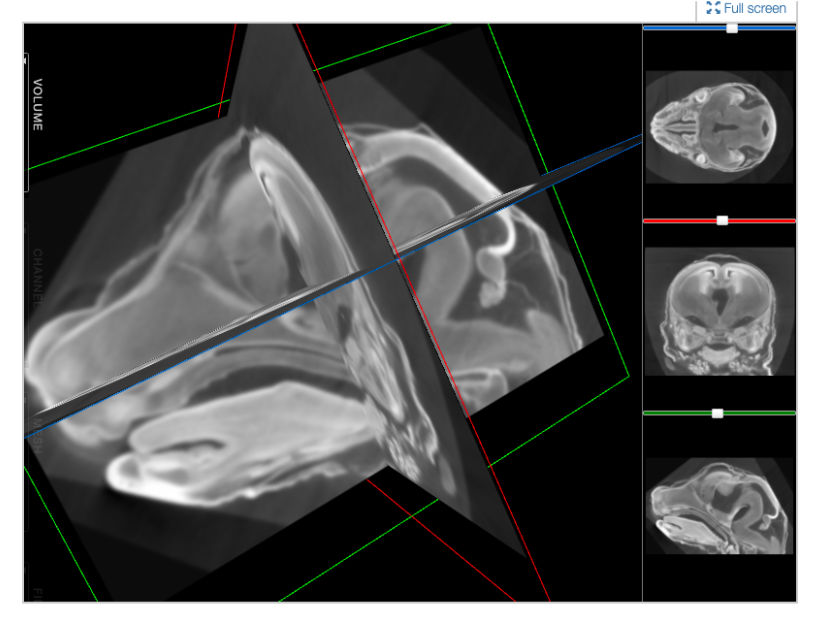
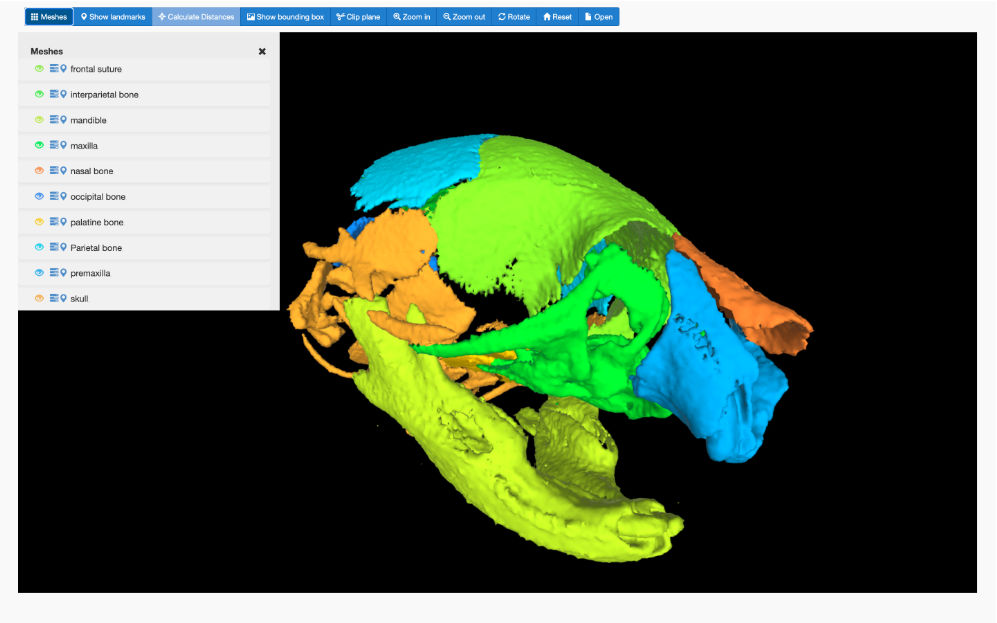
FaceBase is a collaborative NIDCR-funded project that houses comprehensive data in support of advancing research into craniofacial development and malformation. It serves as a community resource by curating large datasets of a variety of types from the craniofacial research community and sharing them via this website. Practices emphasize a comprehensive and multidisciplinary approach to understanding the developmental processes that create the face. The data offered spotlights high-throughput genetic, molecular, biological, imaging and computational techniques. One of the missions of this project is to facilitate cooperation and collaboration between the central coordinating center (ie, the Hub) and the craniofacial research community.
Learn More


GUDMAP
The GenitoUrinary Development Molecular Anatomy Project (GUDMAP) is a consortium of laboratories working to provide the scientific and medical community with tools to facilitate research on the GenitoUrinary (GU) tract. The key components are: a molecular atlas of gene expression for the developing organs of the GU tract; a high resolution molecular anatomy that highlights development of the GU system mouse strains to facilitate developmental and functional studies within the GU system tutorials describing GU organogenesis; and rapid access to primary data via the GUDMAP database
Learn More
RBK
FChronic Kidney Disease (CKD) and Acute Kidney Injury (AKI) pose a substantial public health burden. Even with the best available medical therapy, deteriorating kidney function can require renal replacement therapy (dialysis, kidney transplantation), both of which have substantial morbidity and mortality. Developing and implementing strategies to enhance renal repair and promote the generation of new nephrons in the postnatal organ could have a significant impact on the prevalence and progression of kidney disease. The (Re)Building a Kidney consortium's goal is to coordinate and support studies that will result in the ability to generate or repair nephrons that can function within the kidney.
Learn More