Scientific Projects
Protein Data Bank
We are developing the necessary infrastructure to make the integrative structures of biomolecular systems available from the Protein Data Bank (PDB). PDB is the single global repository for three-dimensional structures of biological macromolecules and their complexes. It is used by millions of scientists and educators worldwide. This project will provide open access to integrative structures through the PDB, which will greatly enhance the understanding of cellular and molecular biology and speed the progress of biomedical and biotechnology research.
Learn More

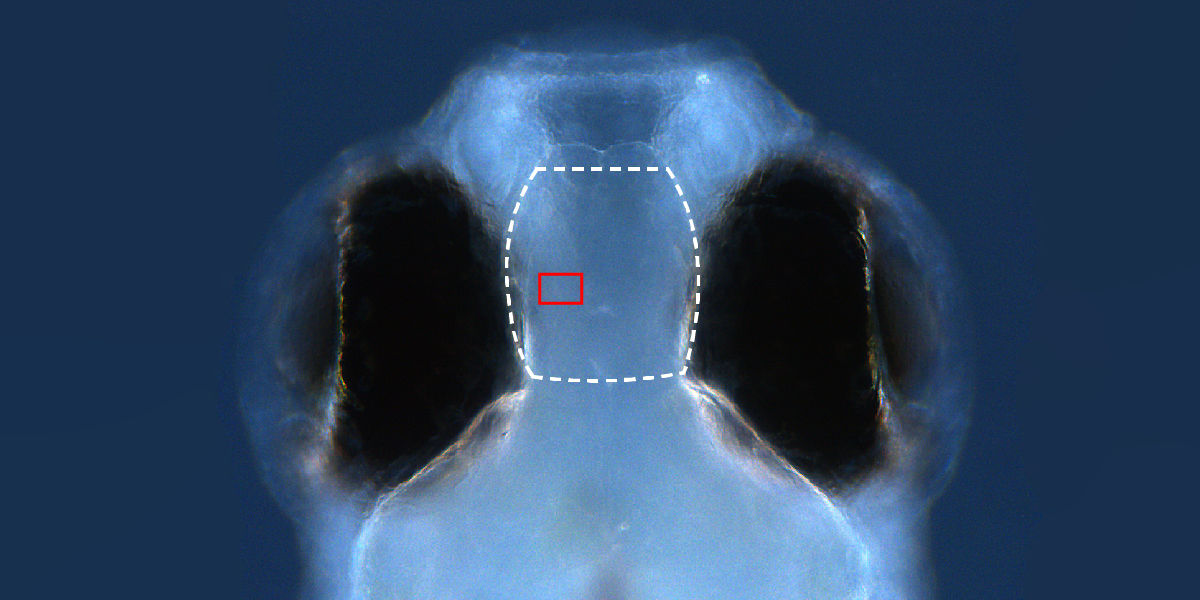
Synapse
We have developed and applied a set of tools that enables direct observation and spatial mapping of synapses using high-contrast selective plane illumination imaging of an endogenous postsynaptic density protein labeled with a recombinant GFP-coupled probe. Using these tools, we have observed changes in the microstructure of the pallium of larval zebrafish that occur during memory formation as a result of exposure to a novel classical conditioning paradigm that we developed. Surprisingly, we found that memory formation due to classical conditioning is associated with generation of new synapses, but not with systematic changes in the strengths of existing synapses.
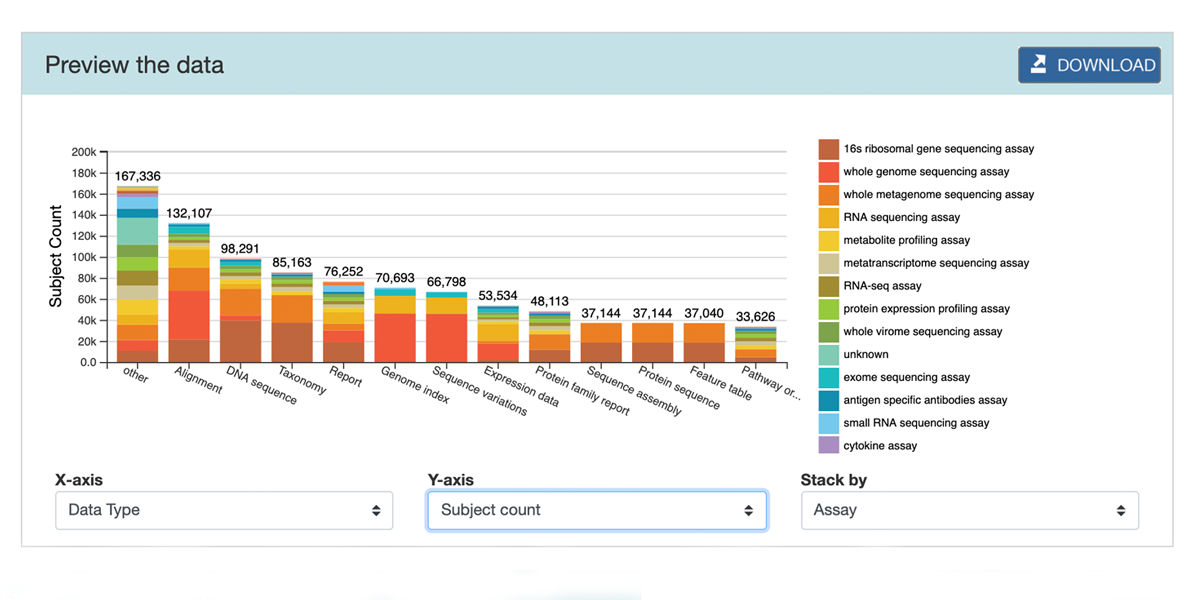
Common Fund Data Ecosystem (CFDE)
NIH Data Commons, a shared virtual space where scientists can work with the digital objects of biomedical research making data-gathering more effective, reliable and accessible The CFDE is an online portal being developed to allow researchers access data from multiple programs in the cloud. ISR is working with CFDE to leverage DERIVA, a cloud-based system they have developed, to help scientists manage big scientific data as easily as pictures on a cell phone.
Learn More